HPC acceleration of FragGeneScan for faster gene prediction on multicore and clustered systems while preserving result correctness.
Context
Gene prediction is a core problem in computational biology, especially for metagenomic workloads where sequence volume and error rates can make processing expensive. FragGeneScan is accurate for these scenarios, but runtime becomes a practical bottleneck at scale.
What I built
I developed an HPC-oriented version of FragGeneScan focused on accelerating execution in multicore cluster environments. The project combines parallel execution approaches and workflow tooling so researchers can process larger inputs faster without changing expected output behavior.
Technical approach
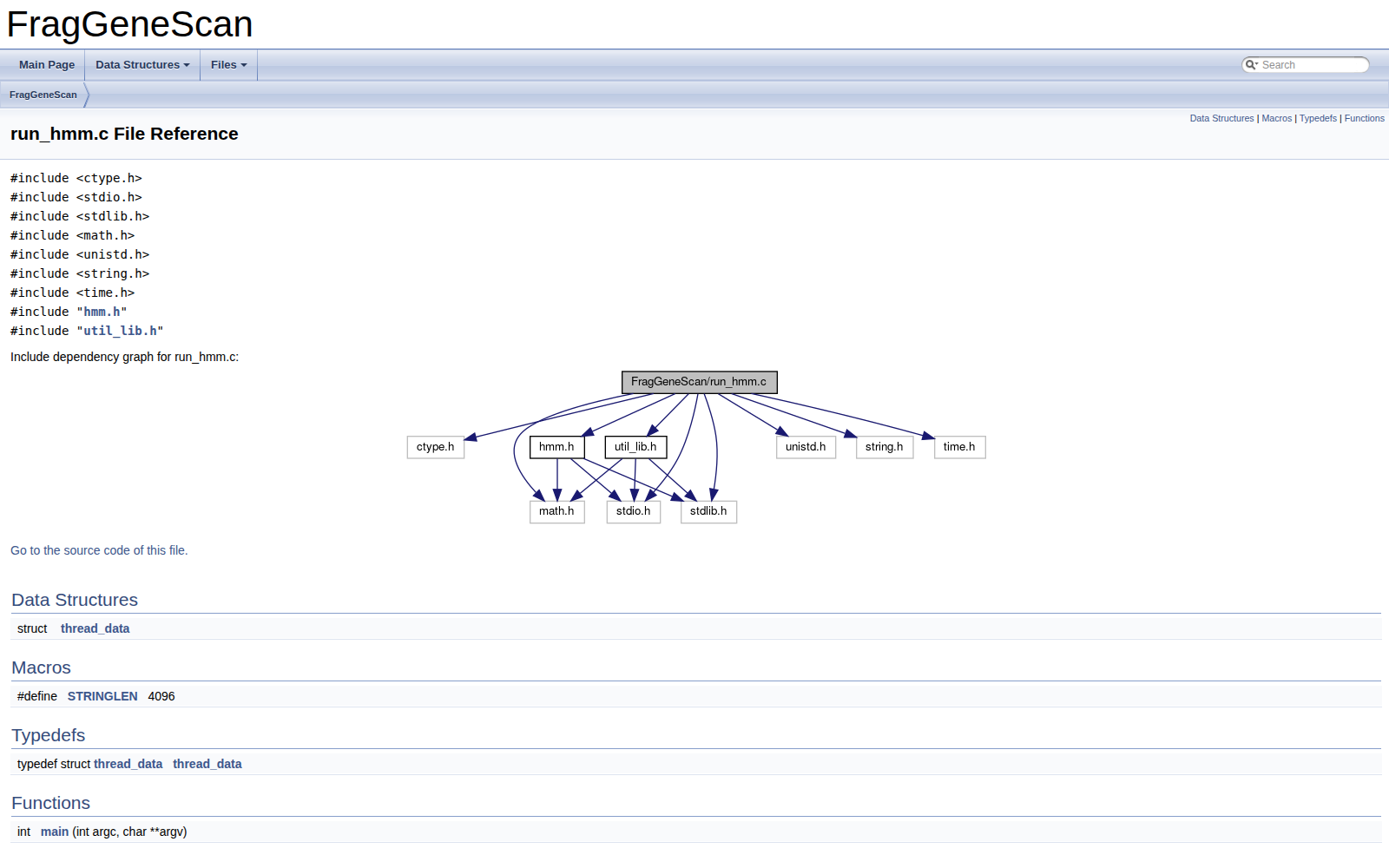
The implementation uses C as the core runtime, with distributed and shared-memory parallelization strategies based on MPI and OpenMP. I also integrated scripts for compilation, execution, and correctness verification to compare results against the reference tool. This keeps performance improvements grounded in reproducibility and scientific validity.
Outcome
FragGeneScan HPC demonstrates that meaningful speedups can be achieved for gene prediction pipelines on multicore clusters while maintaining result consistency with the original method. The project also produced reusable documentation and operational workflows that make the accelerated version easier to adopt in practice.